This package provides access to data frames of values from the COVIDcast
endpoint of the Epidata API. Using the
covidcast_signal() function, you can fetch any data you may
be interested in analyzing, then use
plot.covidcast_signal() to make plots and maps. Since the
data is provided as a simple data frame, you can also wrangle it into
whatever form you need to conduct your desired analyses using other
packages and functions.
Installing
This package is available on CRAN, so the easiest way to install it is simply
install.packages("covidcast")Basic examples
To obtain smoothed estimates of COVID-like illness from our COVID-19 Trends and
Impact Survey for every county in the United States between
2020-05-01 and 2020-05-07, we can use
covidcast_signal():
library(covidcast)
library(dplyr)
cli <- covidcast_signal(data_source = "fb-survey", signal = "smoothed_wcli",
start_day = "2020-05-01", end_day = "2020-05-07",
geo_type = "county")
knitr::kable(head(cli))| data_source | signal | geo_value | time_value | source | geo_type | time_type | issue | lag | missing_value | missing_stderr | missing_sample_size | value | stderr | sample_size |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| fb-survey | smoothed_wcli | 01000 | 2020-05-01 | fb-survey | county | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.8260625 | 0.1341381 | 1676.2773 |
| fb-survey | smoothed_wcli | 01001 | 2020-05-01 | fb-survey | county | day | 2020-09-03 | 125 | 0 | 0 | 0 | 1.0707790 | 0.8213119 | 109.0866 |
| fb-survey | smoothed_wcli | 01003 | 2020-05-01 | fb-survey | county | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.5081644 | 0.2800777 | 572.3194 |
| fb-survey | smoothed_wcli | 01015 | 2020-05-01 | fb-survey | county | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.5277609 | 0.5192431 | 118.8275 |
| fb-survey | smoothed_wcli | 01031 | 2020-05-01 | fb-survey | county | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.3733811 | 0.3367309 | 112.2687 |
| fb-survey | smoothed_wcli | 01045 | 2020-05-01 | fb-survey | county | day | 2020-09-03 | 125 | 0 | 0 | 0 | 1.2369542 | 0.6464530 | 108.5803 |
covidcast_signal() returns a data frame. (Here we’re
using knitr::kable() to make it more readable.) Each row
represents one observation in one county on one day. The county FIPS
code is given in the geo_value column, the date in the
time_value column. Here value is the requested
signal—in this case, the smoothed estimate of the percentage of people
with COVID-like illness, based on the symptom surveys, and
stderr is its standard error. See the
covidcast_signal() documentation for details on the
returned data frame.
To get a basic summary of the returned data frame:
summary(cli)A `covidcast_signal` dataframe with 7030 rows and 15 columns.
data_source : fb-survey
signal : smoothed_wcli
geo_type : county
first date : 2020-05-01
last date : 2020-05-07
median number of geo_values per day : 1015The COVIDcast API makes estimates available at several different
geographic levels, and covidcast_signal() defaults to
requesting county-level data. To request estimates for states instead of
counties, we use the geo_type argument:
cli <- covidcast_signal(data_source = "fb-survey", signal = "smoothed_wcli",
start_day = "2020-05-01", end_day = "2020-05-07",
geo_type = "state")
knitr::kable(head(cli))| data_source | signal | geo_value | time_value | source | geo_type | time_type | issue | lag | missing_value | missing_stderr | missing_sample_size | value | stderr | sample_size |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| fb-survey | smoothed_wcli | ak | 2020-05-01 | fb-survey | state | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.3661909 | 0.1469918 | 1560.000 |
| fb-survey | smoothed_wcli | al | 2020-05-01 | fb-survey | state | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.7764020 | 0.1010989 | 7360.237 |
| fb-survey | smoothed_wcli | ar | 2020-05-01 | fb-survey | state | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.7065584 | 0.1051584 | 4781.483 |
| fb-survey | smoothed_wcli | az | 2020-05-01 | fb-survey | state | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.6025853 | 0.0724214 | 10973.073 |
| fb-survey | smoothed_wcli | ca | 2020-05-01 | fb-survey | state | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.4139045 | 0.0306336 | 50482.138 |
| fb-survey | smoothed_wcli | co | 2020-05-01 | fb-survey | state | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.5984794 | 0.0717395 | 9888.894 |
One can also select a specific geographic region by its ID. For example, this is the FIPS code for Allegheny County, Pennsylvania:
cli <- covidcast_signal(data_source = "fb-survey", signal = "smoothed_wcli",
start_day = "2020-05-01", end_day = "2020-05-07",
geo_type = "county", geo_value = "42003")
knitr::kable(head(cli))| data_source | signal | geo_value | time_value | source | geo_type | time_type | issue | lag | missing_value | missing_stderr | missing_sample_size | value | stderr | sample_size |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| fb-survey | smoothed_wcli | 42003 | 2020-05-01 | fb-survey | county | day | 2020-09-03 | 125 | 0 | 0 | 0 | 0.6270520 | 0.2511377 | 2554.564 |
| fb-survey | smoothed_wcli | 42003 | 2020-05-02 | fb-survey | county | day | 2020-09-03 | 124 | 0 | 0 | 0 | 0.6453498 | 0.2599037 | 2509.176 |
| fb-survey | smoothed_wcli | 42003 | 2020-05-03 | fb-survey | county | day | 2020-09-03 | 123 | 0 | 0 | 0 | 0.5523067 | 0.2497662 | 2473.456 |
| fb-survey | smoothed_wcli | 42003 | 2020-05-04 | fb-survey | county | day | 2020-09-03 | 122 | 0 | 0 | 0 | 0.1430772 | 0.0804642 | 2493.730 |
| fb-survey | smoothed_wcli | 42003 | 2020-05-05 | fb-survey | county | day | 2020-09-03 | 121 | 0 | 0 | 0 | 0.1861889 | 0.0960907 | 2415.204 |
| fb-survey | smoothed_wcli | 42003 | 2020-05-06 | fb-survey | county | day | 2020-09-03 | 120 | 0 | 0 | 0 | 0.3124150 | 0.1218194 | 2465.422 |
API keys
By default, this package submits queries to the API anonymously. All
the examples in the package documentation are compatible with anonymous
use of the API, but there
are some limits on anonymous queries, including rate limits on the
number of queries that can be submitted per hour. To lift these limits,
see the “API keys” section of the covidcast_signal()
documentation for information on how to register for and use an API
key.
Plotting and mapping
This package provides convenient functions for plotting and mapping these signals. For example, simple line charts are easy to construct:
plot(cli, plot_type = "line",
title = "Survey results in Allegheny County, PA")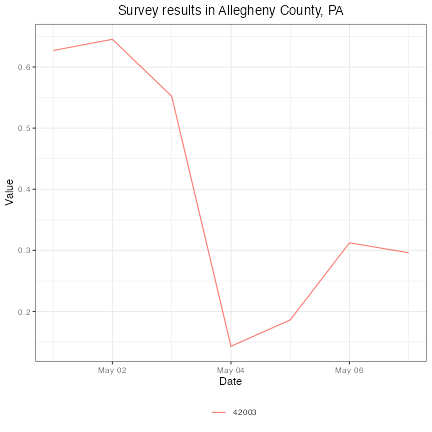
For more details and examples, including choropleth and bubble maps,
see vignette("plotting-signals").
Finding signals of interest
Above we used data from Delphi’s symptom surveys, but the COVIDcast API includes numerous data streams: medical claims data, cases and deaths, mobility, and many others; new signals are added regularly. This can make it a challenge to find the data stream that you are most interested in.
The COVIDcast
Data Sources and Signals documentation lists all data sources and
signals available through COVIDcast. When you find a signal of interest,
get the data source name (such as jhu-csse or
fb-survey) and the signal name (such as
confirmed_incidence_num or smoothed_wcli).
These are provided as arguments to covidcast_signal() to
request the data you want.
Finding counties and metro areas
The COVIDcast API identifies counties by their 5-digit FIPS code and metropolitan areas by their CBSA ID number. (See the geographic coding documentation for details.) This means that to query a specific county or metropolitan area, we must have some way to quickly find its identifier.
This package includes several utilities intended to make the process
easier. For example, if we look at ?county_census, we find
that the package provides census data (such as population) on every
county in the United States, including its FIPS code. Similarly, by
looking at ?msa_census we can find data about metropolitan
statistical areas, their corresponding CBSA IDs, and recent census
data.
(Note: the msa_census data includes types of area beyond
metropolitan statistical areas, including micropolitan statistical
areas. The LSAD column identifies the type of each area.
The COVIDcast API only provides estimates for metropolitan statistical
areas, not for their divisions or for micropolitan areas.)
Building on these datasets, the convenience functions
name_to_fips() and name_to_cbsa() conduct
grep()-based searching of county or metropolitan area names
to find FIPS or CBSA codes, respectively:
name_to_fips("Allegheny")Allegheny County
"42003"
name_to_cbsa("Pittsburgh")Pittsburgh, PA
"38300" Since these functions return vectors of IDs, we can use them to
construct the geo_values argument to
covidcast_signal() to select specific regions to query.
You may also want to convert FIPS codes or CBSA IDs back to
well-known names, for instance to report in tables or graphics. The
package provides inverse mappings county_fips_to_name() and
cbsa_to_name() that work in the analogous way:
county_fips_to_name("42003") 42003
"Allegheny County"
cbsa_to_name("38300") 38300
"Pittsburgh, PA" See their documentation for more details (for example, the options for handling matches when counties have the same name).
Signal metadata
If we are interested in exploring the available signals and their
metadata, we can use covidcast_meta() to fetch a data frame
of the available signals:
meta <- covidcast_meta()
knitr::kable(head(meta))| data_source | signal | time_type | geo_type | min_time | max_time | num_locations | min_value | max_value | mean_value | stdev_value | last_update | max_issue | min_lag | max_lag |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| chng | smoothed_adj_outpatient_cli | day | county | 2020-02-01 | 2023-02-14 | 3118 | 0.0009331 | 99.92012 | 2.227852 | 3.843579 | 1683566979 | 2023-02-19 | 3 | 674 |
| chng | smoothed_adj_outpatient_cli | day | hhs | 2020-02-01 | 2023-06-14 | 10 | 0.0061953 | 20.77577 | 2.530157 | 2.531306 | 1687231582 | 2023-06-19 | 5 | 674 |
| chng | smoothed_adj_outpatient_cli | day | hrr | 2020-02-01 | 2023-06-14 | 306 | 0.0010292 | 50.81590 | 2.350355 | 2.763442 | 1687231582 | 2023-06-19 | 5 | 674 |
| chng | smoothed_adj_outpatient_cli | day | msa | 2020-02-01 | 2023-06-14 | 392 | 0.0007662 | 99.99898 | 2.153204 | 3.000248 | 1687231583 | 2023-06-19 | 5 | 674 |
| chng | smoothed_adj_outpatient_cli | day | nation | 2020-02-01 | 2023-06-14 | 1 | 0.0154639 | 12.08697 | 2.778260 | 2.344107 | 1687231583 | 2023-06-19 | 5 | 674 |
| chng | smoothed_adj_outpatient_cli | day | state | 2020-02-01 | 2023-06-14 | 55 | 0.0013343 | 33.23859 | 2.264207 | 2.563880 | 1687231583 | 2023-06-19 | 5 | 674 |
The covidcast_meta() documentation describes the columns
and their meanings. The metadata data frame can be filtered and sliced
as desired to obtain information about signals of interest. To get a
basic summary of the metadata:
summary(meta)(We silenced the evaluation because the output of
summary() here is still quite long.)
Tracking issues and updates
The COVIDcast API records not just each signal’s estimate for a given location on a given day, but also when that estimate was made, and all updates to that estimate.
For example, consider using our doctor
visits signal, which estimates the percentage of outpatient doctor
visits that are COVID-related, and consider a result row with
time_value 2020-05-01 for geo_values = "pa".
This is an estimate for the percentage in Pennsylvania on May 1, 2020.
That estimate was issued on May 5, 2020, the delay being due to
the aggregation of data by our source and the time taken by the
COVIDcast API to ingest the data provided. Later, the estimate for May
1st could be updated, perhaps because additional visit data from May 1st
arrived at our source and was reported to us. This constitutes a new
issue of the data.
Data known “as of” a specific date
By default, covidcast_signal() fetches the most recent
issue available. This is the best option for users who simply want to
graph the latest data or construct dashboards. But if we are interested
in knowing when data was reported, we can request specific data
versions using the as_of, issues, or
lag arguments. (Note these are mutually exclusive; only one
can be specified at a time.)
First, we can request the data that was available as of a
specific date, using the as_of argument:
covidcast_signal(data_source = "doctor-visits", signal = "smoothed_adj_cli",
start_day = "2020-05-01", end_day = "2020-05-01",
geo_type = "state", geo_values = "pa", as_of = "2020-05-07")A `covidcast_signal` dataframe with 1 rows and 15 columns.
data_source : doctor-visits
signal : smoothed_adj_cli
geo_type : state
data_source signal geo_value time_value source geo_type
1 doctor-visits smoothed_adj_cli pa 2020-05-01 doctor-visits state
time_type issue lag missing_value missing_stderr missing_sample_size
1 day 2020-05-07 6 0 5 5
value stderr sample_size
1 2.581509 NA NAThis shows that an estimate of about 2.3% was issued on May 7. If we
don’t specify as_of, we get the most recent estimate
available:
covidcast_signal(data_source = "doctor-visits", signal = "smoothed_adj_cli",
start_day = "2020-05-01", end_day = "2020-05-01",
geo_type = "state", geo_values = "pa")A `covidcast_signal` dataframe with 1 rows and 15 columns.
data_source : doctor-visits
signal : smoothed_adj_cli
geo_type : state
data_source signal geo_value time_value source geo_type
1 doctor-visits smoothed_adj_cli pa 2020-05-01 doctor-visits state
time_type issue lag missing_value missing_stderr missing_sample_size
1 day 2020-07-04 64 0 5 5
value stderr sample_size
1 5.973572 NA NANote the substantial change in the estimate, to over 5%, reflecting new data that became available after May 7 about visits occurring on May 1. This illustrates the importance of issue date tracking, particularly for forecasting tasks. To backtest a forecasting model on past data, it is important to use the data that would have been available at the time, not data that arrived much later.
Multiple issues of observations
By using the issues argument, we can request all issues
in a certain time period:
covidcast_signal(data_source = "doctor-visits", signal = "smoothed_adj_cli",
start_day = "2020-05-01", end_day = "2020-05-01",
geo_type = "state", geo_values = "pa",
issues = c("2020-05-01", "2020-05-15")) %>%
knitr::kable()| data_source | signal | geo_value | time_value | source | geo_type | time_type | issue | lag | missing_value | missing_stderr | missing_sample_size | value | stderr | sample_size |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-07 | 6 | 0 | 5 | 5 | 2.581509 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-08 | 7 | 0 | 5 | 5 | 3.278896 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-09 | 8 | 0 | 5 | 5 | 3.321781 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-12 | 11 | 0 | 5 | 5 | 3.588683 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-13 | 12 | 0 | 5 | 5 | 3.631978 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-14 | 13 | 0 | 5 | 5 | 3.658009 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-15 | 14 | 0 | 5 | 5 | 3.662286 | NA | NA |
This estimate was clearly updated many times as new data for May 1st arrived. Note that these results include only data issued or updated between 2020-05-01 and 2020-05-15. If a value was first reported on 2020-04-15, and never updated, a query for issues between 2020-05-01 and 2020-05-15 will not include that value among its results.
After fetching multiple issues of data, we can use the
latest_issue() or earliest_issue() functions
to subset the data frame to view only the latest or earliest issue of
each observation.
Observations issued with a specific lag
Finally, we can use the lag argument to request only
data reported with a certain lag. For example, requesting a lag of 7
days means to request only issues 7 days after the corresponding
time_value:
covidcast_signal(data_source = "doctor-visits", signal = "smoothed_adj_cli",
start_day = "2020-05-01", end_day = "2020-05-07",
geo_type = "state", geo_values = "pa", lag = 7) %>%
knitr::kable()Warning: Data not fetched for the following days: 2020-05-01| data_source | signal | geo_value | time_value | source | geo_type | time_type | issue | lag | missing_value | missing_stderr | missing_sample_size | value | stderr | sample_size |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| doctor-visits | smoothed_adj_cli | pa | 2020-05-01 | doctor-visits | state | day | 2020-05-08 | 7 | 0 | 5 | 5 | 3.278896 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-02 | doctor-visits | state | day | 2020-05-09 | 7 | 0 | 5 | 5 | 3.225292 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-05 | doctor-visits | state | day | 2020-05-12 | 7 | 0 | 5 | 5 | 2.779908 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-06 | doctor-visits | state | day | 2020-05-13 | 7 | 0 | 5 | 5 | 2.557698 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-07 | doctor-visits | state | day | 2020-05-14 | 7 | 0 | 5 | 5 | 2.191677 | NA | NA |
Note that though this query requested all values between 2020-05-01 and 2020-05-07, May 3rd and May 4th were not included in the results set. This is because the query will only include a result for May 3rd if a value were issued on May 10th (a 7-day lag), but in fact the value was not updated on that day:
covidcast_signal(data_source = "doctor-visits", signal = "smoothed_adj_cli",
start_day = "2020-05-03", end_day = "2020-05-03",
geo_type = "state", geo_values = "pa",
issues = c("2020-05-09", "2020-05-15")) %>%
knitr::kable()| data_source | signal | geo_value | time_value | source | geo_type | time_type | issue | lag | missing_value | missing_stderr | missing_sample_size | value | stderr | sample_size |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| doctor-visits | smoothed_adj_cli | pa | 2020-05-03 | doctor-visits | state | day | 2020-05-09 | 6 | 0 | 5 | 5 | 2.788618 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-03 | doctor-visits | state | day | 2020-05-12 | 9 | 0 | 5 | 5 | 3.015368 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-03 | doctor-visits | state | day | 2020-05-13 | 10 | 0 | 5 | 5 | 3.039310 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-03 | doctor-visits | state | day | 2020-05-14 | 11 | 0 | 5 | 5 | 3.021245 | NA | NA |
| doctor-visits | smoothed_adj_cli | pa | 2020-05-03 | doctor-visits | state | day | 2020-05-15 | 12 | 0 | 5 | 5 | 3.048725 | NA | NA |